python运行hhsearch二进制命令的包装器类
hhsearch 是 HMM-HMM(Hidden Markov Model to Hidden Markov Model)比对方法的一部分,属于 HMMER 软件套件。它用于进行蛋白质序列的高效比对,特别适用于检测远缘同源性。
以下是 hhsearch 的一些主要特点和用途:
-
HMM-HMM比对:
hhsearch使用隐藏马尔可夫模型(HMM)来表示蛋白质家族的模型。与传统的序列-序列比对方法不同,HMM-HMM比对考虑了氨基酸残基的多序列信息,使得在比对中能够更好地捕捉蛋白质家族的模式和结构。 -
检测远缘同源性:
hhsearch的一个主要优势是其能够检测到相对远离的同源关系。它在比对中引入了更多的信息,从而提高了对远缘同源蛋白的发现能力。 -
灵敏度和特异性:
hhsearch的设计旨在在维持高灵敏度的同时,减少假阳性的比对。这使得它在寻找结构和功能相似性时更为可靠。 -
数据库搜索: 用户可以使用
hhsearch在大型蛋白质数据库中搜索与给定蛋白质序列相似的蛋白质。
"""Library to run HHsearch from Python."""import glob
import os
import subprocess
from typing import Sequence, Optional, List, Iterable
from absl import logging
import contextlib
import tempfile
import dataclasses
import contextlib
import time
import shutil
import re@contextlib.contextmanager
def timing(msg: str):logging.info('Started %s', msg)tic = time.time()yieldtoc = time.time()logging.info('Finished %s in %.3f seconds', msg, toc - tic)@dataclasses.dataclass(frozen=True)
class TemplateHit:"""Class representing a template hit."""index: intname: straligned_cols: intsum_probs: Optional[float]query: strhit_sequence: strindices_query: List[int]indices_hit: List[int]@contextlib.contextmanager
def tmpdir_manager(base_dir: Optional[str] = None):"""Context manager that deletes a temporary directory on exit."""tmpdir = tempfile.mkdtemp(dir=base_dir)try:yield tmpdirfinally:shutil.rmtree(tmpdir, ignore_errors=True)def parse_hhr(hhr_string: str) -> Sequence[TemplateHit]:"""Parses the content of an entire HHR file."""lines = hhr_string.splitlines()# Each .hhr file starts with a results table, then has a sequence of hit# "paragraphs", each paragraph starting with a line 'No <hit number>'. We# iterate through each paragraph to parse each hit.block_starts = [i for i, line in enumerate(lines) if line.startswith('No ')]hits = []if block_starts:block_starts.append(len(lines)) # Add the end of the final block.for i in range(len(block_starts) - 1):hits.append(_parse_hhr_hit(lines[block_starts[i]:block_starts[i + 1]]))return hitsdef _parse_hhr_hit(detailed_lines: Sequence[str]) -> TemplateHit:"""Parses the detailed HMM HMM comparison section for a single Hit.This works on .hhr files generated from both HHBlits and HHSearch.Args:detailed_lines: A list of lines from a single comparison section between 2sequences (which each have their own HMM's)Returns:A dictionary with the information from that detailed comparison sectionRaises:RuntimeError: If a certain line cannot be processed"""# Parse first 2 lines.number_of_hit = int(detailed_lines[0].split()[-1])name_hit = detailed_lines[1][1:]# Parse the summary line.pattern = ('Probab=(.*)[\t ]*E-value=(.*)[\t ]*Score=(.*)[\t ]*Aligned_cols=(.*)[\t'' ]*Identities=(.*)%[\t ]*Similarity=(.*)[\t ]*Sum_probs=(.*)[\t '']*Template_Neff=(.*)')match = re.match(pattern, detailed_lines[2])if match is None:raise RuntimeError('Could not parse section: %s. Expected this: \n%s to contain summary.' %(detailed_lines, detailed_lines[2]))(_, _, _, aligned_cols, _, _, sum_probs, _) = [float(x)for x in match.groups()]# The next section reads the detailed comparisons. These are in a 'human# readable' format which has a fixed length. The strategy employed is to# assume that each block starts with the query sequence line, and to parse# that with a regexp in order to deduce the fixed length used for that block.query = ''hit_sequence = ''indices_query = []indices_hit = []length_block = Nonefor line in detailed_lines[3:]:# Parse the query sequence lineif (line.startswith('Q ') and not line.startswith('Q ss_dssp') andnot line.startswith('Q ss_pred') andnot line.startswith('Q Consensus')):# Thus the first 17 characters must be 'Q <query_name> ', and we can parse# everything after that.# start sequence end total_sequence_lengthpatt = r'[\t ]*([0-9]*) ([A-Z-]*)[\t ]*([0-9]*) \([0-9]*\)'groups = _get_hhr_line_regex_groups(patt, line[17:])# Get the length of the parsed block using the start and finish indices,# and ensure it is the same as the actual block length.start = int(groups[0]) - 1 # Make index zero based.delta_query = groups[1]end = int(groups[2])num_insertions = len([x for x in delta_query if x == '-'])length_block = end - start + num_insertionsassert length_block == len(delta_query)# Update the query sequence and indices list.query += delta_query_update_hhr_residue_indices_list(delta_query, start, indices_query)elif line.startswith('T '):# Parse the hit sequence.if (not line.startswith('T ss_dssp') andnot line.startswith('T ss_pred') andnot line.startswith('T Consensus')):# Thus the first 17 characters must be 'T <hit_name> ', and we can# parse everything after that.# start sequence end total_sequence_lengthpatt = r'[\t ]*([0-9]*) ([A-Z-]*)[\t ]*[0-9]* \([0-9]*\)'groups = _get_hhr_line_regex_groups(patt, line[17:])start = int(groups[0]) - 1 # Make index zero based.delta_hit_sequence = groups[1]assert length_block == len(delta_hit_sequence)# Update the hit sequence and indices list.hit_sequence += delta_hit_sequence_update_hhr_residue_indices_list(delta_hit_sequence, start, indices_hit)return TemplateHit(index=number_of_hit,name=name_hit,aligned_cols=int(aligned_cols),sum_probs=sum_probs,query=query,hit_sequence=hit_sequence,indices_query=indices_query,indices_hit=indices_hit,)def _get_hhr_line_regex_groups(regex_pattern: str, line: str) -> Sequence[Optional[str]]:match = re.match(regex_pattern, line)if match is None:raise RuntimeError(f'Could not parse query line {line}')return match.groups()def _update_hhr_residue_indices_list(sequence: str, start_index: int, indices_list: List[int]):"""Computes the relative indices for each residue with respect to the original sequence."""counter = start_indexfor symbol in sequence:if symbol == '-':indices_list.append(-1)else:indices_list.append(counter)counter += 1class HHSearch:"""Python wrapper of the HHsearch binary."""def __init__(self,*,binary_path: str,databases: Sequence[str],maxseq: int = 1_000_000):"""Initializes the Python HHsearch wrapper.Args:binary_path: The path to the HHsearch executable.databases: A sequence of HHsearch database paths. This should be thecommon prefix for the database files (i.e. up to but not including_hhm.ffindex etc.)maxseq: The maximum number of rows in an input alignment. Note that thisparameter is only supported in HHBlits version 3.1 and higher.Raises:RuntimeError: If HHsearch binary not found within the path."""self.binary_path = binary_pathself.databases = databasesself.maxseq = maxseq#for database_path in self.databases:# if not glob.glob(database_path + '_*'):# logging.error('Could not find HHsearch database %s', database_path)# raise ValueError(f'Could not find HHsearch database {database_path}')@propertydef output_format(self) -> str:return 'hhr'@propertydef input_format(self) -> str:return 'a3m'def query(self, a3m: str) -> str:"""Queries the database using HHsearch using a given a3m."""with tmpdir_manager() as query_tmp_dir:input_path = os.path.join(query_tmp_dir, 'query.a3m')hhr_path = os.path.join(query_tmp_dir, 'output.hhr')with open(input_path, 'w') as f:f.write(a3m)db_cmd = []for db_path in self.databases:db_cmd.append('-d')db_cmd.append(db_path)cmd = [self.binary_path,'-i', input_path,'-o', hhr_path,'-maxseq', str(self.maxseq)] + db_cmdprint("cmd:",cmd)logging.info('Launching subprocess "%s"', ' '.join(cmd))process = subprocess.Popen(cmd, stdout=subprocess.PIPE, stderr=subprocess.PIPE)with timing('HHsearch query'):stdout, stderr = process.communicate()retcode = process.wait()if retcode:# Stderr is truncated to prevent proto size errors in Beam.raise RuntimeError('HHSearch failed:\nstdout:\n%s\n\nstderr:\n%s\n' % (stdout.decode('utf-8'), stderr[:100_000].decode('utf-8')))with open(hhr_path) as f:hhr = f.read()return hhrdef get_template_hits(self,output_string: str,input_sequence: str) -> Sequence[TemplateHit]:"""Gets parsed template hits from the raw string output by the tool."""del input_sequence # Used by hmmseach but not needed for hhsearch.return parse_hhr(output_string)def convert_stockholm_to_a3m (stockholm_format: str,max_sequences: Optional[int] = None,remove_first_row_gaps: bool = True) -> str:"""Converts MSA in Stockholm format to the A3M format."""descriptions = {}sequences = {}reached_max_sequences = Falsefor line in stockholm_format.splitlines():reached_max_sequences = max_sequences and len(sequences) >= max_sequencesif line.strip() and not line.startswith(('#', '//')):# Ignore blank lines, markup and end symbols - remainder are alignment# sequence parts.seqname, aligned_seq = line.split(maxsplit=1)if seqname not in sequences:if reached_max_sequences:continuesequences[seqname] = ''sequences[seqname] += aligned_seqfor line in stockholm_format.splitlines():if line[:4] == '#=GS':# Description row - example format is:# #=GS UniRef90_Q9H5Z4/4-78 DE [subseq from] cDNA: FLJ22755 ...columns = line.split(maxsplit=3)seqname, feature = columns[1:3]value = columns[3] if len(columns) == 4 else ''if feature != 'DE':continueif reached_max_sequences and seqname not in sequences:continuedescriptions[seqname] = valueif len(descriptions) == len(sequences):break# Convert sto format to a3m line by linea3m_sequences = {}if remove_first_row_gaps:# query_sequence is assumed to be the first sequencequery_sequence = next(iter(sequences.values()))query_non_gaps = [res != '-' for res in query_sequence]for seqname, sto_sequence in sequences.items():# Dots are optional in a3m format and are commonly removed.out_sequence = sto_sequence.replace('.', '')if remove_first_row_gaps:out_sequence = ''.join(_convert_sto_seq_to_a3m(query_non_gaps, out_sequence))a3m_sequences[seqname] = out_sequencefasta_chunks = (f">{k} {descriptions.get(k, '')}\n{a3m_sequences[k]}"for k in a3m_sequences)return '\n'.join(fasta_chunks) + '\n' # Include terminating newlinedef _convert_sto_seq_to_a3m(query_non_gaps: Sequence[bool], sto_seq: str) -> Iterable[str]:for is_query_res_non_gap, sequence_res in zip(query_non_gaps, sto_seq):if is_query_res_non_gap:yield sequence_reselif sequence_res != '-':yield sequence_res.lower()if __name__ == "__main__":### 1. 准备输入数据## 输入序列先通过Jackhmmer多次迭代从uniref90,MGnify数据库搜索同源序列,输出的多序列比对文件(如globins4.sto),转化为a3m格式后,再通过hhsearch从pdb数据库中找到同源序列input_fasta_file = '/home/zheng/test/Q94K49.fasta'## input_sequencewith open(input_fasta_file) as f:input_sequence = f.read()test_templates_sto_file = "/home/zheng/test/Q94K49_aln.sto"with open(test_templates_sto_file) as f:test_templates_sto = f.read()## sto格式转a3m格式()test_templates_a3m = convert_stockholm_to_a3m(test_templates_sto)hhsearch_binary_path = "/home/zheng/software/hhsuite-3.3.0-SSE2-Linux/bin/hhsearch"### 2.类实例化# scop70_1.75文件名前缀scop70_database_path = "/home/zheng/database/scop70_1.75_hhsuite3/scop70_1.75"pdb70_database_path = "/home/zheng/database/pdb70_from_mmcif_latest/pdb70"#hhsuite数据库下载地址:https://wwwuser.gwdg.de/~compbiol/data/hhsuite/databases/hhsuite_dbs/ ## 单一数据库#template_searcher = HHSearch(binary_path = hhsearch_binary_path,# databases = [scop70_database_path])## 多个数据库database_lst = [scop70_database_path, pdb70_database_path]template_searcher = HHSearch(binary_path = hhsearch_binary_path,databases = database_lst) ### 3. 同源序列搜索## 搜索结果返回.hhr文件字符串templates_result = template_searcher.query(test_templates_a3m)print(templates_result)## pdb序列信息列表template_hits = template_searcher.get_template_hits(output_string=templates_result, input_sequence=input_sequence)print(template_hits)相关文章:

python运行hhsearch二进制命令的包装器类
hhsearch 是 HMM-HMM(Hidden Markov Model to Hidden Markov Model)比对方法的一部分,属于 HMMER 软件套件。它用于进行蛋白质序列的高效比对,特别适用于检测远缘同源性。 以下是 hhsearch 的一些主要特点和用途: HMM…...

Java 网络编程、e-mail、多线程编程
一、Java 网络编程: 网络编程时指编写运行在多个设备的程序,这些设备通过网络连接起来。 Java.net包中的J2SE的API包含有类和接口,提供低层次的通信细节。 java.net 包中提供了两种常见的网络协议的支持: TCP:TCP&…...

为虚幻引擎开发者准备的Unity指南
目录 1.前言2.编辑器2.1 Scene 视图(视口)2.2 Game 视图 (Play in Editor)2.3.Hierarchy 窗口 (World Outliner)2.4 Project 窗口(Content Browser)2.5 Inspector (Details)2.6 Console(消息视图/输出日志)2.7 Modes 面板在哪里&a…...
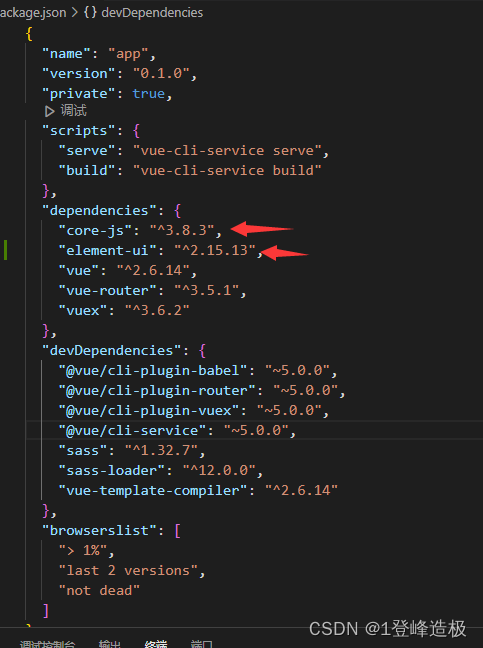
Vue 2使用element ui 表格不显示
直接修改package.json文件 把这两个依赖修改成对应的 删除node_modules 重新安装依赖 重启...

C++学习 --文件
文件操作步骤: 1, 包含头文件#include<fstream> 2, 创建流对象:ofstream ofs 3, 打开文件:ofs.open("文件路径", 打开方式) 4, 写数据:ofs <<…...

java/Android:将字符串按数量分割
分割成数组 import java.util.Arrays;/*** Java将字符串按照指定长度分割成字符串数组*/ public class StringUtils {public static void main(String[] args){String data "227d77a7a244c7b2be3180f2d46be352f56ddf92866692f2cac797358097e5a3e90f6d20bb96bc516a4ab9c0…...

JVM 监控命令详解
文章目录 JDK 中与常用命令行工具jpsjstatjinfojmap导出 dump 文件查看堆内存信息 jstack JVM 可视化分析工具 JDK 中与常用命令行工具 jps 查看当前服务器正在执行的 Java 进程 $> jps 7584 Application 16433 AdminApplication 14209 Jps 5813 Bootstrap 5575 TestApplic…...

TEE威胁评分与评级
目录 一、攻击潜力 1.1 攻击潜力网格 1、实际经过的时间: 2、访问TOE(目标设备)...

-bash: ./deploy.sh: /bin/bash^M: bad interpreter: No such file or directory
文章目录 场景解决 场景 jenkins 发布失败, 报错ERROR: Exception when publishing, exception message [Exec exit status not zero. Status [126]], 这说明远程服务器的deploy.sh执行失败, 首先检查权限,没有发现问题,然后手动执行一遍又报错"-ba…...

【文末送书】十大排序算法C++代码实现
欢迎关注博主 Mindtechnist 或加入【智能科技社区】一起学习和分享Linux、C、C、Python、Matlab,机器人运动控制、多机器人协作,智能优化算法,滤波估计、多传感器信息融合,机器学习,人工智能等相关领域的知识和技术。关…...
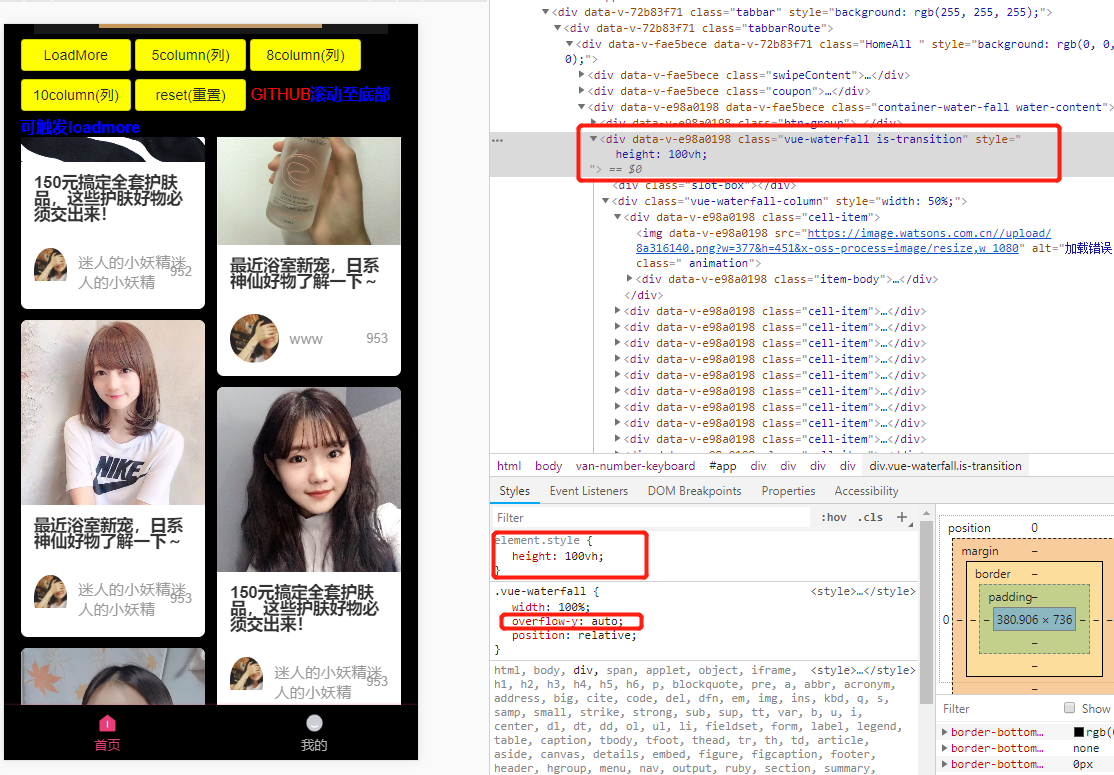
vue-waterfall2 实现瀑布流,及总结的问题
注意:引入需要在主界面引入,直接在组件中引用会有问题 1.安装 npm install vue-waterfall21.8.20 --save (提示:一定要安装1.8.20,最新版会有一部分问题) 2.打开main.js文件 import waterfall from v…...

grafana二次启动失败
背景 安装grafana后启动使用正常,但是关机后再启动显示启动失败,但是看日志又没有报错信息,但是就是启动不了 原因分析 其实是/var/lib/grafana/grafana.db文件损坏了,所以需要把这个文件删掉之后重新启动就正常了,…...

C/C++杂谈-printf的可变参数机制
C/C杂谈-printf的可变参数机制 文章目录 C/C杂谈-printf的可变参数机制printf的使用printf的源码源码剖析 多参数实现机制原理 C11引入了可变参数模板机制,对模板参数进行了高度泛化,但是对于可变参数其实C语言学习中早已遇到过,那就是printf…...
)
es基本语法 (kibana)
#添加 (不添加id默认会生成id) POST /cj/test {"name":"jack","sex":"1","age":12 } #添加 (id为第5个的) POST /cj/test/5 {"name":"jackson","sex":"1","age":12 } #条…...

Tesco EDI需求分析
Tesco,成立于1919年,是一家全球领先的综合性零售企业,总部位于英国。公司致力于提供高质量、多样化的商品和服务,以满足客户的需求。Tesco的使命是通过创新和卓越的客户服务,为客户创造更美好的生活。多年来࿰…...

html常用的标签
基本结构标签 <!DOCTYPE>: 定义 HTML 文档类型。<html>: HTML 文档的根元素。<head>: 文档的头部,包含了元数据和引用的外部资源。<title>: 定义网页标题,显示在浏览器标签上。&l…...

护眼灯什么价位的好?适合学生入手的护眼台灯推荐
据60年前的统计,中国人口的近视率约为10%至20%。 国家卫健委发布的中国首份眼健康白皮书显示,我国小学生近视率为47.2%,初中生近视率为75.8%,大学生近视率超过90%。如今,“低头族”随处可见,近视人群日益增…...

大数据架构
大数据架构 https://huaweicloud.csdn.net/633578fed3efff3090b58398.html https://blog.csdn.net/yuanziok/article/details/117030031 https://blog.csdn.net/qq_46675545/article/details/121985987 https://blog.csdn.net/qq_33367934/article/details/127685417 https://b…...
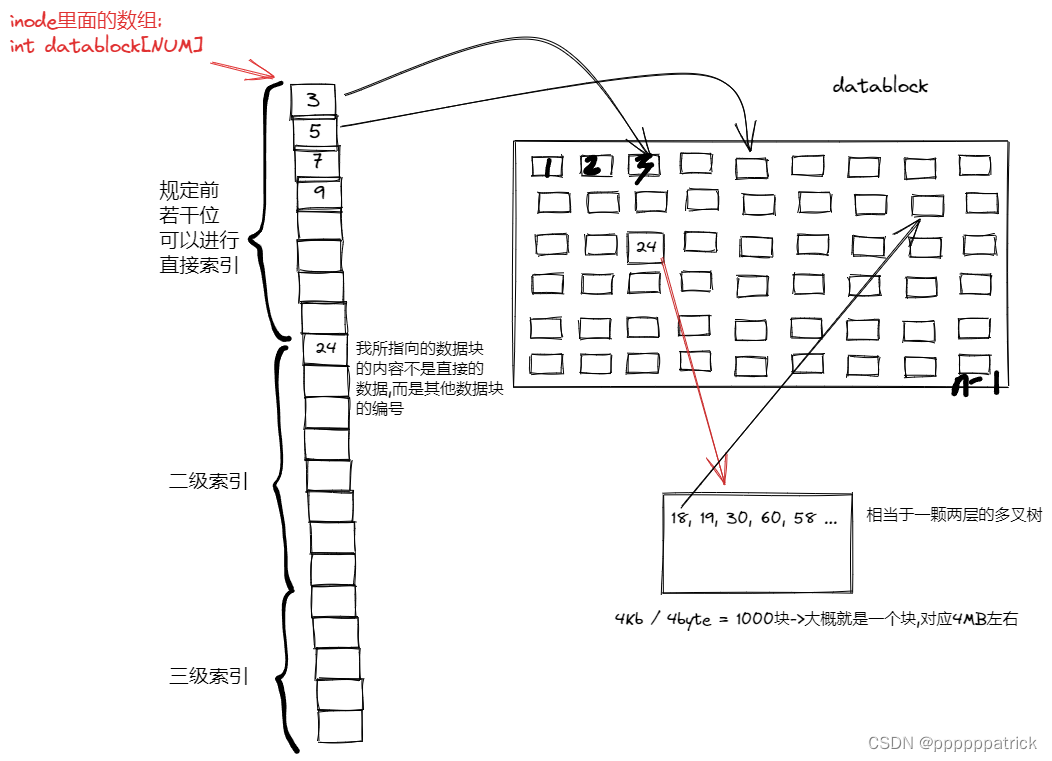
【Linux】C文件系统详解(四)——磁盘的物理和抽象结构
文章目录 磁盘结构磁盘物理结构磁盘的具体物理结构磁盘结构的逻辑抽象 文件系统BootBlockSuperBlockGroupDescriptorTableinode tableDataBlocksinodeBitmapblockBitmaplinux中的inode 和文件名如何理解文件的增删查改删 补充细节1.如果文件误删了,我们该怎么办?2.inode确定分…...

对比自行维护与使用Taotoken在API密钥管理与审计上的差异
🚀 告别海外账号与网络限制!稳定直连全球优质大模型,限时半价接入中。 👉 点击领取海量免费额度 对比自行维护与使用Taotoken在API密钥管理与审计上的差异 在构建基于大模型的应用时,API密钥的管理与审计是保障服务安…...

CANN/ge Format 推导特性分析
Format 推导(Infer Format)特性分析 【免费下载链接】ge GE(Graph Engine)是面向昇腾的图编译器和执行器,提供了计算图优化、多流并行、内存复用和模型下沉等技术手段,加速模型执行效率,减少模型…...

【收藏】2026年AI行业最大的机会,毫无疑问就在应用层!
🔥小白必看|程序员速收藏!2026年AI风口已至,错过再等三年! 字节跳动早已嗅到风口,7个核心团队全速布局Agent智能体,从抖音安全业务到全场景落地,持续加码AI应用层布局;腾…...

CANN ATC模型转换指南
ATC模型转换指南 【免费下载链接】cann-recipes-harmony-infer 本项目为鸿蒙开发者提供基于CANN平台的业务实践案例,方便开发者参考实现端云能力迁移及端侧推理部署。 项目地址: https://gitcode.com/cann/cann-recipes-harmony-infer ATC是异构计算架构CANN…...

SQL PIVOT原理与实战:从行转列到高性能宽表生成
1. 项目概述:从“行变列”开始,真正搞懂SQL PIVOT不是语法糖,而是数据思维的分水岭你有没有遇到过这样的报表需求:销售表里每条记录是一笔订单(客户名、产品名、金额、日期),但老板要的却是“每…...

FPGA-MPSoC边缘AI加速实战:从模型量化到硬件部署全解析
1. 项目概述:为什么要在边缘用FPGA-MPSoC做AI加速?这几年,但凡跟AI沾边的项目,无论是自动驾驶里识别一个突然窜出来的行人,还是工厂质检摄像头判断一个零件的瑕疵,大家挂在嘴边的都是“实时性”和“低功耗”…...

非洲AI本土化实践:医疗、农业、金融、教育四大领域创新与挑战
1. 非洲AI发展的现实图景:机遇与挑战并存 谈论人工智能,我们常常将目光聚焦在硅谷、北京或伦敦。但如果你把视线转向非洲大陆,会发现一片截然不同却又充满生机的AI创新土壤。这里没有OpenAI或DeepMind那样的科技巨头,却有着一群直…...

AI 术语通俗词典:导数
导数是微积分、机器学习、深度学习和人工智能中非常基础的一个术语。它用来描述:当一个输入变量发生微小变化时,函数输出会怎样变化。 换句话说,导数是在回答:如果把输入稍微往前推一点,结果会变大、变小,还…...

基于Claude的智能任务编排引擎:从对话到执行的AI范式跃迁
1. 项目概述:一个基于Claude的智能任务编排与执行引擎最近在GitHub上看到一个挺有意思的项目,叫eyaltoledano/claude-task-master。光看名字,你可能会觉得这又是一个简单的Claude API调用封装。但深入研究后,我发现它的定位远不止…...

AI Agent编排平台ASDM AgentOrbit:从Docker到Kubernetes的生产级部署与管理
1. 项目概述:一个面向生产环境的AI Agent编排与管理平台如果你正在寻找一个能让你像管理服务器一样,轻松创建、部署和管理成百上千个AI Agent实例的平台,那么ASDM AgentOrbit值得你花时间深入了解。这不是一个简单的聊天机器人前端࿰…...

